|
Data Source
For rice, we downloaded the rice cDNA gene set which was provided by TIGR for the NSF rice oligo array project. The FTP URL is here. Certain extra long sequences were broken into pieces to produce the chopped sequences for execution.
Note that these sample runs do not include the mitochondria, chloroplast and transgene gene sets that are included in the NSF funded rice microarray project for their version 2 array design. As such, these oligo sets are not the same as the ones used on that array.
Common Design Parameters
Most design parameters of Picky depend on the experimental conditions. The following design parameters are used to generate the collection of oligos in this page. Except for the maximum_match_len and the minimum_temp_separation parameters, all the other parameters are fixed and their values are shown:
maximum_oligo_size = 70 |
minimum_oligo_size = 50 |
maximum_match_length = 15 |
minimum_match_length = 10 |
maximum_gc_content = 70 |
minimum_gc_content = 30 |
candidates_per_gene = 5 |
probes_per_gene = 1 |
minimum_similarity = 75 |
minimum_temp_separation = 20 |
DNA_concentration = 1 |
Salt_concentration = 75* |
* Note that all Picky reference oligo sets were previously created using Picky 2.0. It is recommended that the new Picky 2.1 be used to redesign any production oligo sets using the hybridization salt concentration from your microarray protocol in order to capture the true benefit of the more precise salt concentration dependant oligo calculation in Picky 2.1.
Variable Design Parameters
Oligo sets can be designed with different stringency by varying the maximum match length and minimum temperature separation parameters of Picky. Generally, a bigger oligo set can be produced when the maximum match is increased, or when the minimum temperature is decreased. However, making those changes also decreased the discriminative power of the oligo designed. Therefore, it is up to the users to decide which oligo set gives them the best balance between number of oligos and the discriminative power. A picture is provided below to illustrate the difference:
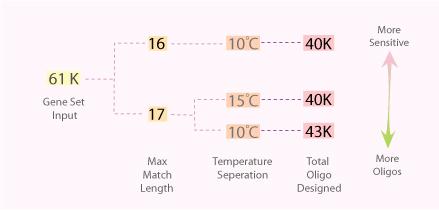
Download Oligo Sets
These oligo sets were computed either on a 2-CPU Opteron 246 2.0Ghz workstation with 4G of total memory, or on a 4-CPU Opteron 848 2.2Ghz server with 32G of memory. Both systems are running Redhat Linux Enterprises OS.
Total Sequences: 61251 |
Maximum Match Length |
Temperature Separation |
16 |
17 |
15°C |
|
(39101 oligos, 4-CPU, 11h 23m) |
10°C |
(39094 oligos, 2-CPU, 10h 1m) |
(43376 oligos, 4-CPU, 10h 33m) |
|