|
Extraction and Conversion of mRNA Coordinates
We will now extract the mRNA coordinates in the Arabidopsis file. In Vect, make sure you are in the 'Input Data' panel with the AC006439.txt file opened. The mRNA coordinates are located next to the mRNA block and are labeled by 'join (' tag (See image below). This selection becomes a little more advanced because in some instances, the 'join(' tag is nested next to a 'complement(' tag. In this example we will be using the New Block Open & Close Condition and Position Independent.
Right click on the 'mRNA' letters and choose New Block Open Condition from the pull down menu. A green box will appear.
Right click on the 'gene' letters and choose New Block Close Condition from the pull down menu . A red box will appear.
Right click and highlight the 'join(' tag and select New Block Open Condition
Then select New Block Close Condition for the ending bracket . Finally, right click on the 'join(' and ' ) ' and select Position Independent from the pull down menu . (You probably have to set Position Independence for 'join(' first before setting the end bracket to Position Independent).
Your data should now look like the diagram below. Position Independent allows the selected tags to be located anywhere in the selection, as opposed to a certain field.
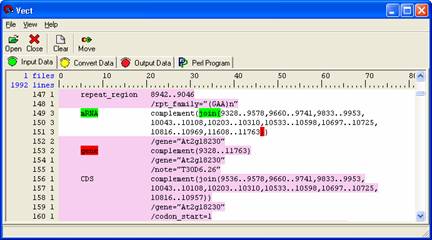
Select the 'Move' icon in the icon panel to move the data to the ‘Convert Data’ panel.
In the 'Convert Data' panel select 'Insert' from the icon panel and apply the 'Filter' rule. Give the rule a descriptive name and select the data from which to apply the rule. Left click on the 'everything' block and select 'integer numbers' from the pull down menu. You now have a list of the raw coordinates.
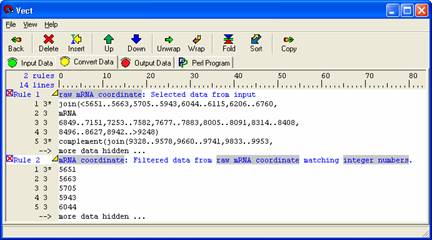
Select Rule 2 (the concatenated sequence ) then click the 'Copy' button from the icon panel to move your data to the 'Output Data' panel. In the 'Output Panel' users can add any text format to the data set and view the changes by selecting the 'Output' icon in the icon panel.
The tag should not be modified but can be moved around. If users wish to limit the output to a set line width, the tag may be edited by including a ':width' before the closing bracket (>). This restricts the body from flowing past the specified width. Example: <gene sequence:60>.
To show the Perl code, move to the 'Perl Program' panel and select 'Compile.' Your Perl program appears as shown below. To run the program generated, select the 'Run' icon. A new window will appear with the results of your Perl program.
If you want to see examples, please check previous mini tutorial for gene sequence extraction and conversion.
 
|